De Novo Protein Sequencing: Basic Concepts and Applications
De novo protein sequencing refers to determining the amino acid sequence of a protein directly without relying on known protein databases. Unlike other analytical methods that depend on known protein sequence databases or known mass spectrometry databases,de novo sequencing utilizes tandem mass spectrometry to directly analyze peptide fragments based on their characteristics. Therefore, peptide sequences not included in protein databases, protein sequences from new species, and protein sequences from genomes that have not yet been sequenced can all be analyzed.Software developed for this method includes PepNovo, PEAKS, and pNovo.
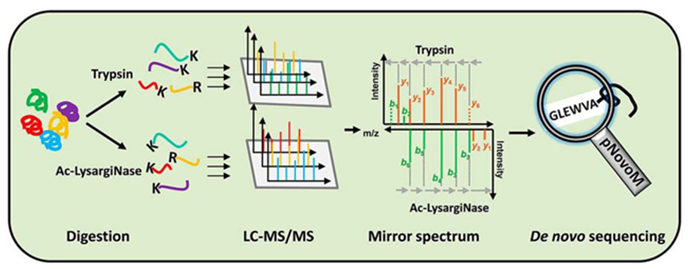
Figure 1. Process of peptide de novo sequencing by mass spectrometry
Basic principles of de novo sequencing
In mass spectrometry, the particles of the substance to be analyzed are ionized before passing through an electromagnetic field. Due toions with different mass-to-charge ratios experiencing different forces in the electromagnetic field, their trajectories and flight times, which allow for the separation and detection of ion characteristics, also differ.During the process of peptide identification using tandem mass spectrometry, de novo sequencing can directly infer the most probable peptide sequence from the secondary spectra. Upon finding specific fragmentation patterns, the corresponding amino acid information can be calculated based on the mass differences between mass spectrometry peaks and post-translational modifications of amino acids.
Applications of de novo sequencing
- Analysis and detection of unknown peptide sequences in biological samples,
- Detection of peptides with missing terminal residues,
- Identification of lysine and leucine,
- Identification of N-terminal blocked and cyclic proteins,
- Accurate determination of sequence information for unknown proteins or peptides to facilitate analysis,
- Functional study of unknown proteins,
- Accurate determination of sequences of commercially modified proteins and enzymes,
- Accurate determination of protein sequences expressed in stable cell lines,
- Monoclonal analysis of antibody primary structure sequences, etc.
De novo protein sequencing is more complex and challenging compared to database-dependent sequencing methods. Short peptide fragments are relatively easy to sequence, but as the fragment length increases, the possible combinations of amino acids increase exponentially, thus increasing the complexity of resolution.
Biotech Co., Ltd.--A premium service provider for bioproduct characterization and multi-omics mass spectrometry detection
Related services:
Protein de novo sequencing and mutation analysis
Monoclonal antibody de novo sequencing
Mass spectrometry-based sequence analysis
Edman degradation-based protein N-terminal sequencing
PCR amplification-based monoclonal antibody sequencing service
How to order?






