Application and Process of Mass Spectrometry Technology in Protein Identification
Mass spectrometry (MS) is a powerful and precise tool for identifying proteins in biological and biochemical research. It allows researchers to identify proteins and their modifications in complex biological samples. Here are the basic steps for protein identification using mass spectrometry:
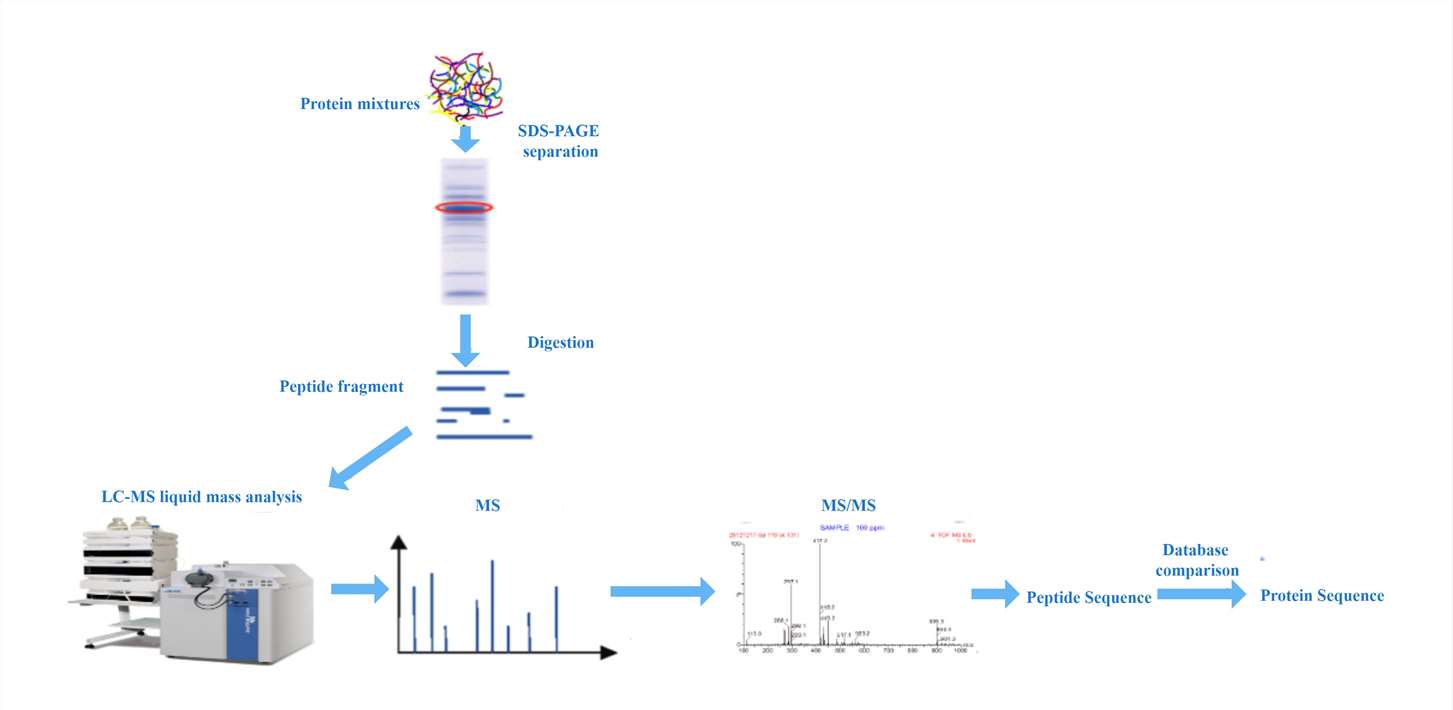
Figure 1. Mass spectrometry sequencing workflow
1. Sample Preparation:
First, proteins need to be extracted from the biological sample and purified using various methods such as centrifugation, filtration, or chromatography. The protein samples are typically digested with an enzyme, commonly trypsin, to cut the proteins into smaller peptides for mass spectrometry analysis.
2. Peptide Separation:
The mixture of digested peptides is separated using techniques like liquid chromatography (LC) or capillary electrophoresis (CE). This step reduces sample complexity, enabling the mass spectrometer to detect each component more effectively.
3. Mass Spectrometry Analysis:
The separated peptides are introduced into the mass spectrometer for analysis. Inside the mass spectrometer, peptides are first ionized (common ionization methods include electrospray ionization (ESI) and matrix-assisted laser desorption/ionization (MALDI)) to generate charged ions. These ions are then separated in the mass analyzer of the spectrometer based on their mass-to-charge ratio (m/z). A mass spectrum is generated by recording the intensity of ions at different m/z values.
4. Data Analysis and Protein Identification:
Information from the mass spectrum is used for protein identification. This is typically done by searching a protein database, matching the experimental data with theoretical mass spectra of known proteins or peptides in the database. Software tools such as Mascot, SEQUEST, or MaxQuant can automate this process to identify the protein composition in the sample.
5. Validation and Quantification:
After identifying specific proteins, further experiments may be needed to validate these findings and quantify the proteins using isotope labeling, label-free quantification methods like TMT, iTRAQ, or SILAC, or label-free approaches such as SWATH.

Figure 1. Example of protein identification based on mass spectrometry by Biotage
The application range of mass spectrometry in protein identification is vast, extending beyond basic protein composition analysis to include but not limited to the following areas:
- Proteomics research: Used for comprehensive analysis of protein expression, composition, and interaction networks in biological samples.
- Protein modification analysis: Mass spectrometry can identify and quantify post-translational modifications (PTMs) of proteins, such as phosphorylation, ubiquitination, methylation, etc.
- Protein structure analysis: Mass spectrometry can provide primary structure information of proteins or peptides, which can be used to infer secondary and tertiary structure characteristics. Cross-linking mass spectrometry (XL-MS) in particular can provide information on interaction sites within protein complexes, aiding in the understanding of protein spatial structure and functional relationships.
- Protein interaction analysis: Mass spectrometry can be used to study protein-protein, protein-DNA, protein-RNA, and protein-small molecule interactions. By analyzing mass spectrometry results after co-immunoprecipitation (Co-IP), the components of protein complexes can be identified, providing key information for studying protein functions and signaling pathways.
- Targeted protein quantitative analysis: Mass spectrometry, particularly multiple reaction monitoring (MRM) technology, can be used for high-sensitivity, high-accuracy protein quantification. This is valuable for biomarker discovery, disease diagnosis, and therapeutic effect evaluation.
- Protein databases and bioinformatics: The analysis of mass spectrometry data relies on protein databases and bioinformatics tools. These tools not only help in identifying unknown proteins but also in protein family classification, functional prediction, and evolutionary analysis.
Through these applications, mass spectrometry has greatly advanced life sciences research, playing an irreplaceable role in areas such as disease mechanisms, drug development, and clinical diagnostics.
BiotechPack, A Biopharmaceutical Characterization and Multi-Omics Mass Spectrometry (MS) Services Provider
Related Services:
Protein mass spectrometry identification
Protein molecular weight determination
Protein identification for gel spots, gel strips, and IP samples
Peptide mass spectrometry identification
Shotgun protein identification
Membrane protein identification service
Pull-down target protein mass spectrometry identification
Protein interaction mass spectrometry analysis
Post-translational modification proteomics analysis
N/C-terminal protein sequencing
Comprehensive protein analysis
How to order?






